 Natasha, Ana, Katie, and Johnson (from L to R) Natasha, Ana, Katie, and Johnson (from L to R) Ana Vasquez-a rising senior at South El Monte High School in Los Angeles- participated in the Upward Bound program at Harvey Mudd this summer. She spent four weeks with us learning the ins-and-outs of DNA extraction, PCR, and gel electrophoresis. She worked on samples from the NOAA Restore Act. We collected several unidentified octocorals, and she was able to amplify mtMutS and COI for them. This helped us to put tentative names on these unidentified octocorals (Villogorgia sp. and Nicella sp.). Ana is planning on attending college, but does not know yet what she wants to major in, although she knows for sure that she would like to minor in music or Spanish. She plays trumpet in marching band at South El Monte. Ana has been in the Harvey Mudd Upward Bound program for almost three years. It was a pleasure to have her in lab this summer!
1 Comment
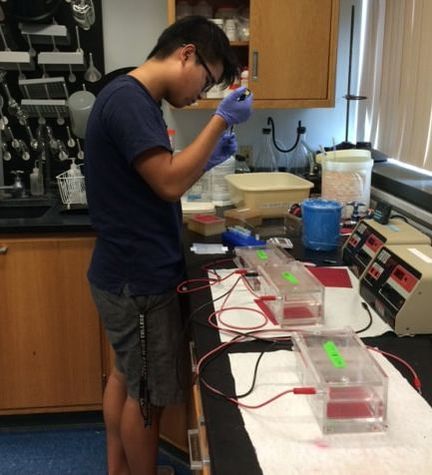
 Katie Erickson (HMC '19) is preparing the remaining Restore samples for sequencing Katie Erickson (HMC '19) is preparing the remaining Restore samples for sequencing With only two weeks of research left for me and many of my student collaborators, we are working on wrapping up our work for the summer to get it ready for presentation and publication. A large portion of this work has consisted of manually fixing and confirming gene annotations of complete mitochondrial genomes. Johnson and I have been working to assemble and annotate the mitochondrial genomes of Octocorallia, while Natasha and Brooks have focused on Hexacorallia. We are finishing up our annotation revisions so we will be submitting our sequences to GenBank soon. We also will be using the protein coding genes that we have annotated to build a phylogenetic tree based on mitochondrial data. Johnson and I will be comparing the tree built off of mitochondrial genes to a tree built from nuclear data as a way to compare the methods and what they tell us about the history and relationships of Octocorallia. I’ve spent much of my time in the lab this summer working on DNA Barcoding for coral samples collected from the Northern Gulf of Mexico. These barcodes contribute to a project funded by the NOAA RESTORE Act that asseses the Population Connectivity of Deepwater Corals in the Northern Gulf of Mexico. Since the target species of the project have all been characterized with DNA barcodes, we can use the sequencing technique as a way to confirm the species of the samples. I have focused on sequencing the mitochondrial genes MutS for Callogorgia samples and COI for Paramuricea biscaya. Thus far, I have confirmed the morphological identification of all of the samples I have sequenced. I have also identified the different haplotypes of Callogorgia and Paramuricea and will continue to analyze how these haplotypes are distributed spatially across the northern Gulf of Mexico. I have also had the opportunity to mentor and work with two high school students in the lab this summer who have helped me with DNA barcoding. Justin has helped me with the barcoding for the target specimens in the RESTORE project, while Ana has sequenced the mitochondrial gene MutS for samples collected from the same locations whose species identifications are unknown. Overall, I am astonished by how much I have learned in one summer, and have a lot of excitement as I analyze the results of our work and get to see how it contributed to larger projects.  HMC '20 student Johnson Hoang posing in front of the Coral Reefs Tank at the Aquarium of the Pacific. Lab field trip!!! HMC '20 student Johnson Hoang posing in front of the Coral Reefs Tank at the Aquarium of the Pacific. Lab field trip!!! It’s only been six weeks since I’ve started working in the McFadden lab, but I’ve been pipetting a lot and learning even more. As some of my other labmates have previously mentioned, one of the projects we are working on is to try resolve the phylogeny of Anthozoa. One of the ways we are doing this is through the analyzing the ultra-conserved elements (UCEs). This begins with library prep of DNA, which I have almost finished for the Alcyonium samples. Alcyonium is a genera of coral that are unique in the sense that they have no calcium carbonate skeleton. Despite the lack of a skeleton, species with Alcyonium are essential to marine habitats. However, the distinction between species is not well understood and so we hope that with the UCEs, we may be able to not only differentiate populations of Alcyonium but be able to delineate species within the genus as well. Hopefully we can soon obtain the reagents we need to finish target and enrichment and start analyzing UCEs because I am excited to see what new evolutionary insights we can learn. On the bioinformatic side of things, I am also going through the process of annotating and cleaning up mitochondrial genomes of several octocorals. As Natasha and Brooks have mentioned, we have taken assemblies from several different programs and annotated them using MITOS 2. While MITOS 2 gives us a pretty good idea of the genes within the mitochondrial genome it isn’t perfect. We have to painstakingly fix contigs ourselves in order to get the proper gene length, codons, etc. However, we are working on a script to try and help automate the process and so I look forward to that. Once that’s done, we’ll be able to construct phylogenetic trees based off of mitochondrial genomes and compare it to other data. So far, majority of the mitochondrial genomes I have analyzed match the ancestral gene order presented in Brockman’s and McFadden’s 2012 paper on mitochondrial genomes with the exception of Protodendron repens, whose classification is within the family Alcyoniidae but mitochondrial genome better matches those within the family Xeniidae. Overall, we’ve made a lot of progress in both in the wet and dry lab. Although our work isn’t done yet, I’m getting very excited for the potential discoveries we may encounter.  Brooks MacDonald-HMC class of 2020-conducting library prep on Sinularia spp. Brooks MacDonald-HMC class of 2020-conducting library prep on Sinularia spp. After showing up a week late to the summer research party, I’ve had the opportunity to jump into the lab and bioinformatics work of the McFadden lab. It’s been a whirlwind learning all the lab techniques and the biology behind the work we’re doing, but I’m gaining a better understanding of the work done in this lab. I’ve also learned a ton about all the prior work done in and out of this lab that has attempted to resolve the phylogeny of Anthozoa. I’ve spent most of my time doing library preparations on the sheared DNA of Sinularia, soft corals dubbed “leather corals.” This is the initial process to ultimately sequence and analyze the ultraconserved elements (UCEs) of their genomes. This week I am finishing up all the Sinularia samples we have, so I am also helping to finish the Actinaria (sea anemone) samples. In the next week or two, we will likely move forward and do targeted enrichment on the DNA libraries in order to prepare for sequencing. After sequencing, we will move forward with bioinformatics approaches, though the summer is likely to be over by the time this round of samples is finished. However, there is other bioinformatics that I’ve been working on. I’ve just finished annotating the mitochondrial genomes of specimens in the hexacorallian order Zoantharia. This process involves taking assemblies done by Trinity, SPAdes, and/or Novoplasty and using MITOS 2 to annotate the genes. I’ve seen a couple differences so far in the presence of introns tRNAs, and genome length--between samples I’ve analyzed this summer and between our samples and the zoanthid mt genomes published in the literature. The mt genomes of Zoantharia have not been very thoroughly examined in the literature, so the work I am doing will help add to the scientific conversation, which is very exciting. It’s been a lot of fun to work with the other students in the lab, and it’s a great experience to be able to work under people with as much knowledge as Prof. McFadden and Dr. Quattrini. I’m looking forward to analyzing some of our results more in depth and thinking about what they mean in the broader context of the phylogeny of anthozoans. I’m almost at the halfway point of my summer research in the McFadden lab. I’m working with three other Mudd students this summer, and each of us is in charge of overseeing a different aspect of our project -- all related to understanding the evolutionary history and connectivity of corals and their relatives the anemones, as well as trying to delimit species, using methods such as barcoding and target enrichment. Since the samples I’m focusing on, the anemones Actinaria, weren’t ready for the first couple of weeks, I helped the other team members out with their projects. These included DNA extractions and barcoding for several Paramuricea samples, as well as library preparation of Alcyonium samples. Last week, the 64 Actinaria samples arrived, so I was able to begin library preparation on them, and I’ll be working on this for most of the next few weeks. The goal of this library preparation is target-capture enrichment of UCE (Ultra-Conserved Element) and exon loci, which will help us determine the species boundaries of closely-related anemone species. This information could help answer questions such as whether different color morphs of the same anemone species actually represent unique species. When I’m not doing library preparation, I’m doing bioinformatics work in the dry lab. With regards to bioinformatics, I’ve mostly been working on annotating the mitochondrial genomes of the hexacoral order Antipatharia (black corals) using the programs MITOS, Spades, and Novoplasty. These annotations will further help us understand the connections between coral species. It’s amazing to have the opportunity to work with such great mentors, and I’m learning a ton.
 Rei Imada (left) and Justin Jiang (right) presenting research on defining operative taxonomic units of Xenia sp. with DNA barcodes. Rei Imada (left) and Justin Jiang (right) presenting research on defining operative taxonomic units of Xenia sp. with DNA barcodes. As school commences for the fall term, Harvey Mudd hosted it's annual poster session yesterday, highlighting summer research projects across the campus. Our students, Mike Adams (HMC '18) and Rei Imada (Claremont-McKenna '20) presented posters on their summer research projects. Mike worked on examining whether invertebrates have obligate associations with corals in the deep Caribbean, Rei presented his research on barcoding xeniid octocorals from shallow waters off Australia. Rei was assisted this summer by Justin Jiang=a current junior at Walnut High School. A lot of progress was made during this summer, and we thank our students for their hard work and dedication to their projects!  Although Shelia is spending her time in the lab this summer learning new techniques, she has previously spent time at sea. Shelia is seen here in the ROV control van on the R/V Manta. Although Shelia is spending her time in the lab this summer learning new techniques, she has previously spent time at sea. Shelia is seen here in the ROV control van on the R/V Manta. As an undergraduate student at New York City College of Technology in Brooklyn, New York, I conducted research on the molecular characterization of Antipatharians (black corals) under the guidance of Professor Mercer R. Brugler and Dr. Estefania Rodriguez in the Black Coral Lab at the American Museum of Natural History. This summer I was fortunate enough to participate in a NSF funded summer research exchange program in Professor McFadden’s lab at Harvey Mudd College in Claremont, California. I’ve been working closely with Dr. Andrea Quattrini and fellow student researcher Alicia Pentico (Harvey Mudd ’19) on the Ultraconserved Elements (UCEs) project in an attempt to clarify phylogenetic relationships within the class Anthozoa. For my portion of the project, I’ve been focusing on relationships among orders of Hexacorallia. The subclass consists of six orders: Actiniaria, Antipatharia, Ceriantharia, Corallimorpharia, Scleractinia and Zoantharia. Over the last nine weeks I’ve split my time pretty evenly between working in the wet lab on library preparation and hybridization and working in the dry lab on bioinformatics. I’ve constructed three phylogenetic trees using three datasets: mitochondrial DNA, nuclear rDNA (18s and28s) and UCEs. I look forward to the next steps of this project and seeing what evolutionary relationships will be revealed once Alicia and I combine our data.  Mike Adams (HMC '18) (on left) is in his PPE on Oceaneering's ship Ocean Project in the Gulf of Mexico. Mike Adams (HMC '18) (on left) is in his PPE on Oceaneering's ship Ocean Project in the Gulf of Mexico. This summer the Anthozoan UCE’s project and Dr. Quattrini have given me the opportunity to take part in my first ever deep-sea research cruise! We are currently on day 7 of the 17-day OP2017 Restore Cruise in the northern Gulf of Mexico. We are targeting 4 species of coral (Hypnogorgia pendula, Paramuricea biscaya, Swiftia exserta, and Callogorgia delta) to sample across 16 sites. The cruise is a collaboration between labs at Harvey Mudd College and Lehigh University and is funded by a grant from the NOAA Restore Act. Populations of the target species were heavily impacted by the Deepwater Horizon oil spill in 2010 and by the subsequent use of chemical dispersants to clean up the oil. The overarching goal of the project is to eventually determine a model of gene flow of these 4 species in the Gulf of Mexico to guide restoration efforts for the impacted populations. The models of gene flow and larval dispersal will be created using the results of population genomics conducted by the Herrera Lab at Lehigh University and DNA barcoding efforts conducted by Dr. Quattrini in the McFadden Lab at Harvey Mudd College. The genetic results combined with larval dispersal models (A. Bracco, GA Tech) will give us an idea about patterns of connectivity between populations across the GOM. My roles on the cruise include both watch standing and sample processing. My sample processing duties involve preserving a clipping of all the coral samples we collect in 95% Ethanol for later DNA barcoding. I am not only doing this for our own barcoding purposes at Harvey Mudd, but I am also preserving mesophotic corals (H. pendula and S. exertia) in ethanol for Dr. Etnoyer’s Lab at NOAA. My watch standing duties include logging dives in the ROV van when the ROV is collecting samples and otherwise aiding anyone in anything they need done while I am on watch. While those are my official duties, they are not the only activities I have been participating in during my stay on the ship. I have been aiding other scientists on board with their projects when they need help and I have learned a lot about other ongoing research in the field of deep-water corals and their associates. Beyond this I have been spending time making lasting connections with great young minds in the field at institutions such as Lehigh, Temple, Penn State, University of Hawaii, NOAA, and USGS. I am very appreciative of this amazing opportunity that has solidified my ambitions of continuing to pursue a career in deep-sea exploration and research. -Mike Adams (Harvey Mudd College ’18)  The three tracks in the photo above are as follows: genome (scf180000022261), annotation from Nematostella (first_try.gff), annotation from Mnemiopsis (whole_pipe.gff). Mnemiopsis predicted one large gene with many introns and exons, but Nematostella predicted many smaller genes. The three tracks in the photo above are as follows: genome (scf180000022261), annotation from Nematostella (first_try.gff), annotation from Mnemiopsis (whole_pipe.gff). Mnemiopsis predicted one large gene with many introns and exons, but Nematostella predicted many smaller genes. For the past month, I have been assembling and annotating the genome of Renilla muelleri, commonly known as a sea pansy. We isolated DNA from organisms acquired through the aquarium trade, which were then sequenced using both an Illumina Hi-Seq, and MiSeq and a Pacific Biosciences RS II. Our paired-end Illumina reads had an average length of 166 bp at 190x coverage, and our PacBio reads were at 10x coverage. I fed the Illumina reads and the PacBio subreads to MaSuRCA-3.2.2 for a hybrid de novo assembly; the resulting scaffold was run through stats.sh (bbmap) and Quast to calculate genome statistics. Both programs reported the genome to be ~185 Mb with a GC content of 36.3%. The assembly had 6,036 contigs with a N50 of 63.191 Kb. To compare different assemblies, we used SPAdes to assemble a genome with only the Illumina reads. Using the same statistics programs, the Illumina-only assembly had a similar GC content 36.9%, but a genome size of ~ 255 Mb. I attribute these differences to the difficulties that short-read-only assemblers have in resolving repeat regions. The assembler is unable to identify the length of a certain region because of numerous base pair repeats unless it has longer reads like PacBio. I then used the draft hybrid assembly for annotation with Augustus, a gene prediction software. I conducted two annotations with Augustus, one using training data from Nematostella vectensis (provided by Joe Ryan) and the other with Mnemiopsis leidyi; both used RNASeq data of Renilla provided by J. Ryan. Based on previous research, the number of genes in Renilla is anywhere from 15,000 to 25,000. With the Nematostella training set, 20,464 genes were predicted, and with Mnemiopsis 8,588 genes were predicted. The Mnemiopsis training set was able to predict exons and introns, but the Nematostella training set seemed to have predicted each exon as a gene. Because of this, the Mnemiopsis annotation had less than half of the predicted genes Nematostella annotation had. I tested these training sets out, but I realize that Nematostella and Mnemiopsisa are >500 MY divergent from Renilla. In the future, I will use a training set specific to Renilla generated from RNA-seq data. I am also planning on assembling a final genome with environmental contaminants (e.g., bacteria, viruses) removed and then running a final annotation. Stay posted for the draft genome and annotation files! Justin Jiang, Walnut High School ‘19 |
UCE Project TeamAll things Anthozoa, Evolution and Ecology Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation
Archives
September 2016
Categories |





 RSS Feed
RSS Feed
